Looking to the Future
In this section you will learn:
- The next generation of technologies that are fueling the full spectrum of cancer science from bench to bedside will accelerate the pace of understanding of cancer biology while transforming the future of clinical practice.
- Combining genomic and proteomic approaches in cancer research will revolutionize treatment by expanding precise use of existing therapeutics and addressing some of the most elusive questions in cancer such as treatment resistance.
- Artificial intelligence holds enormous promise in cancer science and medicine and may transform the future of cancer detection, diagnosis, treatment, and drug discovery.
- The new wave of innovation in science and technology is providing researchers with the necessary tools to effectively target cancer-driving molecules that have long been “undruggable.”
- Implementation science aims to integrate proven, effective interventions into everyday life and routine health care in order to bridge the gap between evidence and clinical practice.
These are unprecedented times for the medical research community. While advances across the spectrum of cancer science and medicine are fueling unparalleled progress against cancer and driving down the U.S. cancer death rates, the medical research community has also faced extraordinary adversity over the past two years in the face of the COVID-19 pandemic. However, many researchers, including AACR President, 2021–2022, David A. Tuveson, MD, PhD, FAACR, are extremely hopeful about the future because they are confident that through innovative and collaborative research, we will power more advances against cancer while we continue to control the public health crisis caused by COVID-19. The new wave of scientific and technological innovations discussed in this chapter has the potential to transform cancer research and patient care in the years to come.
Proteogenomics: A New Frontier in Precision Cancer Medicine
As described in Understanding How Cancer Develops, the order of the four bases in DNA provides the code used by a cell to produce the various proteins it needs to function. The genetic code in the DNA is first converted into another molecule called ribonucleic acid (RNA) which is then used by the cell to manufacture proteins. Once manufactured, proteins undergo various chemical modifications such as receiving carbohydrate or lipid tags leading to the formation of a mature protein. All important functions within a cell are carried out by proteins. One can generally conclude that normal DNA encodes normal proteins, which helps produce normal cells, which assemble into healthy tissues. Conversely, altered DNA leads to altered proteins which may lead to unhealthy tissues and tumor development.
While cancer genomics—the comprehensive analysis of tumor-associated genetic changes—has become a core component of modern precision medicine, looking exclusively at the DNA (or RNA) provides an incomplete picture of the biological underpinnings of cancer etiology. This is because mutational analysis, albeit an important tool, cannot always reliably predict changes in the level or function of the corresponding proteins. Such limitations are highlighted by the fact that while genomic databases, such as The Cancer Genome Atlas, catalog numerous genetic changes associated with multiple cancer types, the impact of many of those mutations on cellular function or on patients’ outcomes remains unknown. It should also be noted that even though the presence of specific genetic alterations is frequently used as a biomarker to determine whether a patient is eligible for treatment with a molecularly targeted therapeutic, the main component of precision medicine, most of these therapeutics work by binding to tumor proteins. Adding proteomics—the comprehensive analysis of all the proteins inside a cell—to the armamentarium of cancer research, can be a powerful tool to gain novel insights into a patient’s tumor that cannot be realized by genomics alone. Researchers strongly believe that when used together, cancer proteomics and genomics can truly open new opportunities in the diagnosis, prognosis, and treatment of cancers and transform the landscape of patient care.
According to NCI, proteogenomics is defined as the study of how information about the DNA in a cell or organism relates to the proteins made by that cell or organism. This includes understanding when proteins get made and what modifications occur to proteins after they are made that may switch them on or off. Ongoing research on cancer proteogenomics, much of which is led by the NCI’s Clinical Proteomic Tumor Analysis Consortium (CPTAC), is aimed at uncovering novel insights into the cellular and molecular basis of cancer development and identifying potential new therapeutic interventions that cannot be obtained through genomics alone (605)Rodriguez H, Zenklusen JC, Staudt LM, Doroshow JH, Lowy DR. The next horizon in precision oncology: Proteogenomics to inform cancer diagnosis and treatment. Cell. 2021;184(7):1661-1670.(606)Chen YJ, Roumeliotis TI, Chang YH, et al. Proteogenomics of Non-smoking Lung Cancer in East Asia Delineates Molecular Signatures of Pathogenesis and Progression. Cell. 2020;182(1).(607)Krug K, Jaehnig EJ, Satpathy S, et al. Proteogenomic Landscape of Breast Cancer Tumorigenesis and Targeted Therapy. Cell. 2020;183(5).(608)Petralia F, Tignor N, Reva B, et al. Integrated Proteogenomic Characterization across Major Histological Types of Pediatric Brain Cancer. Cell. 2020;183(7).(609)Huang C, Chen L, Savage SR, et al. Proteogenomic insights into the biology and treatment of HPV-negative head and neck squamous cell carcinoma. Cancer Cell. 2021;39(3).(610)Satpathy S, Krug K, Beltran PMJ, et al. A proteogenomic portrait of lung squamous cell carcinoma. Cell. 2021;184(16):4348-4371.e40..
Using proteogenomics, researchers can classify tumors based on their molecular characteristics rather than cellular morphologies, which has traditionally been the leading approach for tumor classification. Identification of the molecular characteristics provides key insights to a tumor’s potential therapeutic vulnerabilities. As one example, a comprehensive proteogenomic analysis of more than 100 breast cancer samples performed in a recent study provided an in-depth look at the inappropriate activation of certain cellular pathways and aberrant cellular metabolism within subsets of breast cancers. The researchers identified a subset of patients with hormone receptor-positive breast tumors, who are usually treated with targeted therapeutics, as potential candidates for treatment with immune checkpoint inhibitors, thereby increasing the utility of these revolutionary immunotherapeutics in a new group of cancer patients (607)Krug K, Jaehnig EJ, Satpathy S, et al. Proteogenomic Landscape of Breast Cancer Tumorigenesis and Targeted Therapy. Cell. 2020;183(5).. In a second study, proteogenomic analysis helped researchers understand why certain patients diagnosed with head and neck cancers whose tumors present abnormal levels of EGFR or the immune checkpoint protein PD-L1 still do not respond to EGFR-targeted therapeutics or immune checkpoint inhibitors (609)Huang C, Chen L, Savage SR, et al. Proteogenomic insights into the biology and treatment of HPV-negative head and neck squamous cell carcinoma. Cancer Cell. 2021;39(3).. These data are critical for the selection of appropriate patient populations that are most likely to respond to EGFR-targeted therapies or immunotherapies. In addition, the study identified novel targets that could be used for the development of future checkpoint inhibitors against these aggressive cancers. Yet another recent proteogenomic analysis from more than 200 pediatric brain tumor patients representing seven cancer types offered novel insights into the molecular alterations within brain tumors and revealed previously unknown therapeutic avenues for certain children with brain cancers (608)Petralia F, Tignor N, Reva B, et al. Integrated Proteogenomic Characterization across Major Histological Types of Pediatric Brain Cancer. Cell. 2020;183(7).. The data uncovered a new molecularly targeted treatment option for certain children with a brain tumor known as craniopharyngioma, which currently has no effective therapeutic options.
Novel insights obtained through proteogenomic analysis of cancers have tremendous diagnostic and therapeutic potential and may provide answers to some of cancer’s most elusive questions such as treatment resistance and recurrence. However, the use of proteogenomics is not yet a part of routine clinical practice. Some of the current challenges that researchers are trying to overcome include identifying methodologies that allow for combined analysis of DNA, RNA, and protein from limited tissue samples obtained through biopsies (611)Satpathy S, Jaehnig EJ, Krug K, et al. Microscaled proteogenomic methods for precision oncology. 2020;11(1):1-16.; improving tissue acquisition techniques; and overcoming logistic and workflow barriers to maintain accurate documentation of samples in large-scale research studies (612)Yoo S, Shi Z, Wen B, Kho S, Pan R, Feng H, et al. A community effort to identify and correct mislabeled samples in proteogenomic studies. Patterns 2021;2:100245.. Going forward, we anticipate that the next wave of innovation in science and technology along with advances in cross-disciplinary collaboration, data sharing, and patient engagement will lead to the integration of proteogenomics into the fabric of cancer research and care. This exciting new frontier in precision cancer medicine is poised to transform the future landscape of cancer diagnosis, prognosis, and treatment, bringing new hope to many more patients with cancer.
Artificial Intelligence: Shaping the Future of Cancer Science and Medicine
According to NCI, artificial intelligence (AI) is defined as the ability of a computer to perform functions that are usually thought of as intelligent human behavior, such as learning, reasoning, problem solving, and decision-making. As researchers accumulate large quantities of cancer-related data ranging from tumor images from scans or pathological slides, to tumor sequencing, electronic health records, and clinical outcomes, AI can analyze this information to derive meaningful insights that previously could not have been realized (613)Topol EJ. High-performance medicine: the convergence of human and artificial intelligence. Nat Med 2019 251. 2019;25(1):44-56.). Machine learning is an application of AI that focuses on the development of computer programs that can access and learn from data, identify patterns, and make decisions without explicit human intervention. Deep learning is a subset of machine learning that utilizes neural networks to make decisions. The applications of AI in cancer science and medicine are vast and rapidly expanding. For instance, AI has the potential to streamline processes for radiological and pathological image interpretation allowing for faster decision-making for people with life-threatening cancers. Several clinical applications of AI in radiology and pathology were discussed in the AACR Cancer Progress Report 2020 (467)Sengupta R, Honey K. AACR Cancer Progress Report 2020: Turning Science into Lifesaving Care. Clin Cancer Res. 2020;26(19).. Some recent advances in the field that are already benefiting patients are described in previous chapters in this report (see Enhancing the Speed and Accuracy of Interpreting Cancer Screening Tests and Using Artificial Intelligence for Precision in Surgical Oncology). In this section we provide a brief overview of selected exciting new AI approaches that are currently in investigation and may provide future breakthroughs in cancer care.
Transforming Drug Development
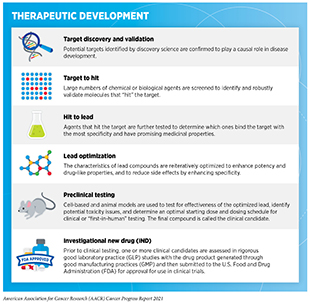
Researchers are harnessing the power of AI in different ways to accelerate drug discovery for many diseases including cancer (614)Ho D. Artificial intelligence in cancer therapy. Science (80- ). 2020;367(6481).(615)Sahoo D, Swanson L, Sayed IM, et al. Artificial intelligence guided discovery of a barrier-protective therapy in inflammatory bowel disease. Nat Commun. 2021;12(1).(616)Savage N. Tapping into the drug discovery potential of AI. Biopharma Deal. Published online 2021.. Some efforts are aimed at accelerating the pace of basic research such as identifying new targets, while others are designed to make clinical trials more efficient. The use of AI can potentially improve each step of the drug development process (see sidebar on Therapeutic Development). For instance, AI can harness massive amounts of information from the scientific literature, clinical databases, and patient-derived data to identify potential new drug targets, such as proteins that are vital for cancer growth; to design new therapeutics that target such proteins; and to help evaluate the safety and effectiveness of those therapeutics (617)J V, D C, P C, et al. Applications of machine learning in drug discovery and development. Nat Rev Drug Discov. 2019;18(6):463-477..
As mentioned in the discussion of proteogenomics, most molecularly targeted therapeutics that are an integral part of precision medicine target tumor-associated proteins in cancer cells or in the tumor microenvironment. To design therapeutics that can effectively attach to and modulate protein function, researchers must know the 3-dimensional structure of proteins. Traditionally, solving protein 3-dimensional structures has been difficult and time-consuming since it requires complex experiments, including crystallization of proteins followed by visualization using X-rays and highly sophisticated electron microscopes. Recently, an AI program called AlphaFold was shown to be able to predict 3-dimensional structures of proteins with incredible precision and accuracy (354)Callaway E. “It will change everything”: DeepMind’s AI makes gigantic leap in solving protein structures. Nature 2020;588:203–4.(618)Senior AW, Evans R, Jumper J, et al. Improved protein structure prediction using potentials from deep learning. 2020;577(7792):706-710.(619)Service RF. ‘The game has changed.’ AI triumphs at solving protein structures. Science 2020;370:1144–5.. In fact, the program outperformed around 100 research teams in a prestigious biennial protein-structure prediction challenge called Critical Assessment of Structure Prediction. Researchers have further used AlphaFold to predict the structures of almost all 20,000 proteins that are expressed by the human genome as well as many others belonging to other organisms (620)Tunyasuvunakool K, Adler J, Wu Z, et al. Highly accurate protein structure prediction for the human proteome. Nature. Published online 2021.. Scientists anticipate that tools such as AlphaFold will help drug development researchers predict the 3-dimensional structures of proteins of interest rapidly and economically. They are confident that AlphaFold as well as other similar AI platforms will revolutionize the future of drug development for complex diseases such as cancer by significantly decreasing the time and costs associated with traditional approaches (620)Tunyasuvunakool K, Adler J, Wu Z, et al. Highly accurate protein structure prediction for the human proteome. Nature. Published online 2021.(621)Jumper J, Evans R, Pritzel A, et al. Highly accurate protein structure prediction with AlphaFold. Nature. Published online 2021..
Decades of basic research have taught us that immune cells called T cells are naturally capable of destroying cancer cells. We have also learned that tumors evade destruction by T cells because they have high levels of proteins that attach to and trigger “brakes” on T cells, stopping the T cells from attacking the tumor (see Releasing the Brakes on the Immune System). These brakes, which are proteins on the surface of T cells, are called immune-checkpoint proteins. It took researchers over a decade since the discovery of the first checkpoint inhibitor to develop therapeutics that target these proteins. Checkpoint inhibitors have revolutionized cancer treatment, even for patients with very advanced cancers. Notably, the impact of AI in accelerating immune checkpoint drug development is evident from recent research. As one example, an AI platform was able to discover a potential checkpoint inhibitor candidate within only eight months (622)Exscientia announces first AI-designed immuno-oncology drug to enter clinical trials. [cited 2021 Jul 10].. The therapeutic blocks the activation of the checkpoint protein adenosine 2a receptor on T cells, for which there are no currently available inhibitors approved by FDA. Based on promising preclinical data from pancreatic and lung cancer models which show that the therapeutic is able to activate cancer-fighting T cells and reduce the number of viable tumor cells (623)Payne A, Fons P, Alt I, Van Ham J, Taubert C, Bell A, et al. 1731 – EXS21546, a non-CNS penetrant A 2A R-selective antagonist for anti-cancer immunotherapy. Session PO.IM02.07, Immunomodulatory Agents and Interventions. 2021 April 10. AACR Annual Meeting, the candidate molecule will soon be evaluated in phase I clinical trials (624)NIH, U.S. National Library of Mecine, Clinical Trials.gov. 3-part study to assess safety, tolerability, pharmacokinetics and pharmacodynamics of EXS21546. [updated 2021 Jan 29; cited 2021 July 10]..
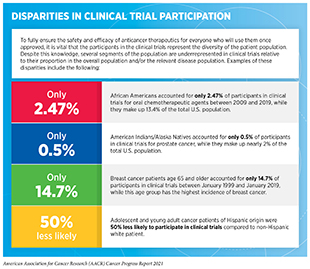
One of the most pressing challenges in drug development is low patient participation in clinical trials, especially for racial and ethnic minorities and other underserved populations (see sidebar on Disparities in Cancer Clinical Trial Participation). Overcoming barriers to clinical trial participation is a major focus for all stakeholders in the cancer research community. Data from recent studies indicate that AI platforms can play a critical role in these efforts. A known barrier for minority patients’ participation in clinical trials is restrictive and sometimes poorly justified eligibility criteria for patient inclusion. A new report, which used an AI platform to harness data from electronic health records from more than 60,000 lung cancer patients and publicly available trial eligibility criteria from clinicaltrials.gov to evaluate the real-world impact of eligibility criteria on patient recruitment and outcomes, found that many patients who were excluded from certain trials due to the restrictive criteria could have benefited from treatments provided in the trials (625)Liu R, Rizzo S, Whipple S, et al. Evaluating eligibility criteria of oncology trials using real-world data and AI. 2021;592(7855):629-633.. In fact, when the researchers broadened the eligibility criteria using the AI-guided approach, the estimated number of eligible patients more than doubled. Notably, the study also found that trials with broader eligibility did not have any more adverse event-related treatment withdrawals compared to trials with strict eligibility criteria. AI-driven methodologies such as the one used in this study can be critical in the future design of more inclusive clinical trials while still maintaining patient safety.
Predicting Patient Outcomes
Across the continuum of cancer care, there is growing interest in utilizing AI to harness patients’ data to guide disease management such as evaluating cancer susceptibility for high-risk individuals, making treatment decisions, and predicting treatment responses and long-term outcomes (626)Manz CR, Chen J, Liu M, et al. Validation of a Machine Learning Algorithm to Predict 180-Day Mortality for Outpatients with Cancer. JAMA Oncol. 2020;6(11).(627)Song J, Wang L, Ng NN, et al. Development and Validation of a Machine Learning Model to Explore Tyrosine Kinase Inhibitor Response in Patients with Stage IV EGFR Variant-Positive Non-Small Cell Lung Cancer. JAMA Netw Open. 2020;3(12).(628)(629)Tseng YJ, Wang HY, Lin TW, Lu JJ, Hsieh CH, Liao CT. Development of a Machine Learning Model for Survival Risk Stratification of Patients with Advanced Oral Cancer. JAMA Netw Open. 2020;3(8):1-15..
Patients with HCV infection-induced chronic cirrhosis have a high risk of developing a form of liver cancer known as hepatocellular carcinoma. According to a recent retrospective analysis, an AI program outperformed standard statistical models at identifying individuals with liver cirrhosis who were likely to develop liver cancer (628)Ioannou GN, Tang W, Beste LA, Tincopa MA, Su GL, Van T, et al. Assessment of a deep learning model to predict hepatocellular carcinoma in patients with hepatitis C cirrhosis. JAMA Netw Open 2020;3:e2015626.. By predicting which individuals had the highest risk of developing cancer the AI program provides avenues for prioritizing patient surveillance and treatment. In a second study, a deep learning-based AI approach was able to use CT images from patients with stage IV NSCLC that contained alterations in the EGFR gene to identify which patients are most likely to benefit from EGFR-targeted therapeutics (627)Song J, Wang L, Ng NN, et al. Development and Validation of a Machine Learning Model to Explore Tyrosine Kinase Inhibitor Response in Patients with Stage IV EGFR Variant-Positive Non-Small Cell Lung Cancer. JAMA Netw Open. 2020;3(12).. These data can help health care providers not only to select patients who are most likely to benefit from therapy, but also to spare those who are unlikely to benefit to avoid unnecessary adverse effects of such treatments. In a third study a machine learning program was able to utilize data from electronic health records to prospectively identify cancer patients who are at high risk of short-term mortality within six months after their encounter with the health care system (626)Manz CR, Chen J, Liu M, et al. Validation of a Machine Learning Algorithm to Predict 180-Day Mortality for Outpatients with Cancer. JAMA Oncol. 2020;6(11).. Such AI-based tools may allow health care providers to inform appropriate behavioral interventions for patients and engage in a timelier conversation regarding the patients’ goals of care and end-of-life preferences.
Collectively, these studies emphasize the incredible potential of AI for the future of clinical cancer care. However, one area where researchers must pay close attention is the inclusion of datasets from diverse populations that are representative of the overall U.S. cancer patient population during the development of AI platforms. Lack of diversity in the data that are used to develop AI or machine learning systems may incorporate racial/ethnic or other biases within AI applications and limit their generalizability for different patient population groups who must benefit from these state-of-the-art technologies. As one example, since many of the AI programs aimed to detect skin cancers were trained primarily on light-skinned individuals, there are concerns that these tools perform poorly in detecting skin cancer affecting individuals with darker skin (630)Adamson AS, Smith A. Machine learning and health care disparities in dermatology. JAMA Dermatology. 2018;154(11).. Similar concerns have been raised by experts in cancer genomics research based on many recent findings demonstrating that samples used in cancer genomics research projects such as cancer-related genome-wide association studies are collected primarily from white populations (631)Gao Y, Cui Y. Deep transfer learning for reducing health care disparities arising from biomedical data inequality. Nat Commun. 2020;11(1).. Addressing these challenges will require both short- and long-term approaches ranging from more diverse data collection and AI program monitoring to infrastructural changes such as diversification of the funders and developers of AI research, publication, and education (632)Zou J, Schiebinger L. Ensuring that biomedical AI benefits diverse populations. EBioMedicine. 2021;67..
New Wave of Innovation to Aim at Cancer’s Most Intractable Targets
Since the signing of the National Cancer Act 50 years ago, there has been unprecedented progress in the treatment of the collection of diseases we call cancer. The newest pillars of cancer treatment, molecularly targeted therapy and immunotherapy, which form the foundation of precision medicine, have revolutionized care for many patients leading to remarkable durable responses even in individuals with metastatic cancer. Notably, a majority of these cutting-edge cancer treatments are small molecules or antibodies which work by physically binding to cellular proteins in tumors and blocking their function. Unfortunately, targeted therapeutics and immunotherapeutics are only available for a selected number of patients with certain types of cancers and even among patients with cancers that are targetable with molecularly targeted therapeutics or immunotherapeutics, most patients ultimately develop resistance to these drugs.
Even though numerous genes and their encoded proteins have been shown to be altered in cancers, one of the key challenges in precision medicine is that scientists, so far, have only been able to successfully target a fraction of these molecules therapeutically. For instance, while it has been long recognized that mutations in TP53, RAS, or MYC genes can promote tumor growth and are very common in many cancers, the respective proteins have been difficult to target. The key challenges that have prevented researchers from developing therapeutics against some of these proteins include their location within a cell and their structure. Proteins localized on the surface of a cell are easier to target compared to those that are inside a cell. In addition, it is easier for scientists to design therapeutics against proteins that have ordered 3-dimensional structures and defined binding pockets which can serve as efficient docking sites for the therapeutics. Often these binding pockets are adjacent to the active sites, which are regions of the protein that are critical for their function. In fact, many proteins that have been difficult to target, e.g., p53, RAS, or MYC, are localized inside the cell and possess highly disordered and labile 3-dimensional structures. Powered by a new wave of scientific and technological innovations, such as those described here, researchers are now looking into exciting new approaches to target some of the most intractable proteins involved in cancer development.
Flagging Cancer Cells for Destruction by the Immune System
Immunotherapeutics unleash a patient’s own immune system to fight cancer and can work in different ways. One approach that researchers have utilized for the development of immunotherapeutics is designing molecules that flag cancer cells in some way for detection and destruction by the immune system (see Unleashing the Body’s Defense System Against Cancer). Antibodies are a class of immune system proteins that are frequently used to flag cancer cells. In two recent studies, researchers were able to target two of the most elusive proteins associated with cancer, RAS and p53 (633)Hsiue EH-CC, Wright KM, Douglass J, et al. Targeting a neoantigen derived from a common TP53 mutation. Science (80- ). 2021;371(6533).(634)Douglass J, Hsiue EH-CC, Mog BJ, et al. Bispecific antibodies targeting mutant RAS neoantigens. Sci Immunol. 2021;6(57):5515., using engineered antibodies. Mutations in RAS and p53 have been detected in 50 percent and 30 percent of all tumors, respectively, and although there has been some recent progress in treating cancers driven by RAS mutations (see A New Breakthrough in Treating Lung Cancer), researchers are still looking for ways to target p53. Targeting p53 mutations has been particularly challenging since in many cases the mutant form of the protein is inactive, and since most cancer drugs work by blocking the function of overactive proteins rather than restoring the function of inactive ones. In addition, both RAS and p53 are located inside the cell making therapeutic targeting of these molecules especially difficult.
Decades of basic research in cancer immunology, coupled with the latest advances in molecular biology, proteomics, and bioinformatics tools, have enabled researchers to overcome some of the challenges in drug targeting. It is now known that cells display fragments of intracellular proteins on their surface during a process that is needed for normal immune function. In the case of cancer cells, some of these fragments are parts of mutated proteins which are specific only to cancer cells but absent in normal healthy tissue. By using this knowledge, researchers were able to design bispecific antibodies that attached to fragments of either mutated p53 or RAS on the cancer cell surface with one end and to immune cells with the other end. By bringing immune cells close to cancer cells the antibodies help immune cells to recognize and eliminate cancer cells. In preclinical studies, these antibodies were able to slow tumor growth. These data highlight an exciting new approach in targeting difficult to reach proteins, and future research will determine whether these novel antibodies can improve outcomes for patients with cancer harboring mutations in p53 or RAS.
Mobilizing Targeted Protein Degradation
The proteasome is a molecular machine naturally found in cells that breaks down proteins the cell no longer needs. The process helps control multiple functions including cell division and survival. Selective degradation of cancer-causing proteins using the proteasome machinery is an approach that is currently being tested, especially for proteins that have been difficult to target by conventional methods (635)Samarasinghe KTG, Crews CM. Targeted protein degradation: a promise for undruggable proteins. Cell Chem Biol. Published online 2021.. One area of active investigation is the development of Proteolysis Targeting Chimeras (PROTACs), a class of therapeutics that can induce targeted degradation of disease-causing proteins (635)Samarasinghe KTG, Crews CM. Targeted protein degradation: a promise for undruggable proteins. Cell Chem Biol. Published online 2021.(636)Dang C V., Reddy EP, Shokat KM, Soucek L. Drugging the “undruggable” Cancer Targets. Vol 17. Nature Publishing Group; 2017:502-508.(637)Coleman N, Rodon J. Taking aim at the undruggable. Am Soc Clin Oncol Educ Book 2021;41:1–8.. These bifunctional small molecules consist of two protein-binding elements that are attached by a linker; one binds to the protein of interest (target) and another recruits an E3 ubiquitin ligase, a key component of the proteasomal machinery. By bringing the target close to the E3 ligase, PROTACs initiate breakdown and elimination of the target proteins.
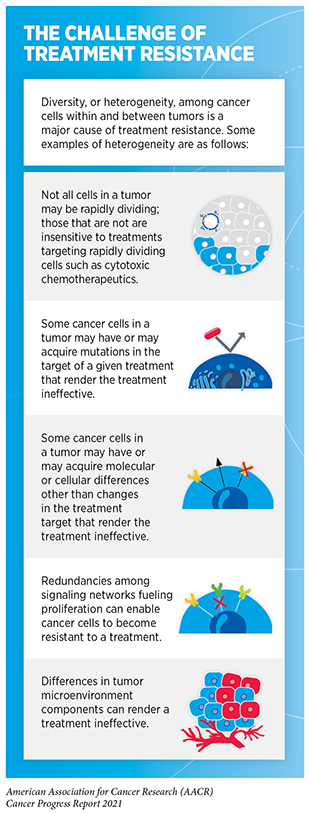
Because of their unique mechanism of action, PROTACs have certain advantages over traditional cancer treatment using small molecule inhibitors or antibodies (e.g., most molecularly targeted therapeutics or immunotherapeutics) which work by physically binding to cellular proteins in tumors and blocking their function. An ongoing challenge in treatment with molecularly targeted therapeutics or immunotherapeutics is the development of treatment resistance (see sidebar on The Challenge of Treatment Resistance). Common mechanisms by which cancer cells become resistant to therapeutics are by increasing the levels of target proteins or by gaining new mutations in the target proteins. Since PROTACs work by destroying rather than inhibiting target proteins, treatment with PROTACs can overcome these pathways of treatment resistance. Another advantage with this type of therapeutics approach is that PROTACs are recycled after they degrade their target proteins. Therefore, one molecule of PROTAC can eradicate multiple copies of the target protein. Thus, a smaller dose of PROTACs compared to small molecule inhibitors might be as effective in producing a desired antitumor effect. Furthermore, since PROTACs do not need to bind to the active sites of target proteins, they can be easier to design against otherwise intractable targets.
PROTACs targeting a wide range of targets for many cancer types are currently in different phases of preclinical and clinical development (638)Mullard A. Targeted protein degraders crowd into the clinic. Nat Rev Drug Discov 2021;20:247–50.. Among these are efforts to use PROTACs to degrade otherwise difficult to target cancer-causing proteins such as p53, STAT3, RAS, MYC, etc. (638)Mullard A. Targeted protein degraders crowd into the clinic. Nat Rev Drug Discov 2021;20:247–50.(639)Bery N, Miller A, Rabbitts T. A potent KRAS macromolecule degrader specifically targeting tumours with mutant KRAS. 2020;11(1):1-14.(640)Wang C, Zhang J, Yin J, et al. Alternative Approaches to Target Myc for Cancer Treatment. Vol 6. Nature Publishing Group; 2021:1-14.(641)Hines J, Lartigue S, Dong H, Qian Y, Crews CM. MDM2-recruiting PROTAC offers superior, synergistic antiproliferative activity via simultaneous degradation of BRD4 and stabilization of p53. Cancer Res. 2019;79(1).. To maximize their therapeutic function, researchers are exploring ways to activate PROTACs in tumor-specific manners as well as in selected tissues. In this regard, there are ongoing efforts aimed at developing modified versions of PROTACs such as those that can be activated by irradiation or attachment to tumor-specific ligands (e.g., Antibody Conjugated Bifunctional Degraders) (635)Samarasinghe KTG, Crews CM. Targeted protein degradation: a promise for undruggable proteins. Cell Chem Biol. Published online 2021.. Targeted protein degradation using PROTACs, as well as the newer generation of more sophisticated protein degraders, holds great promise for the future of cancer medicine and may transform cancer treatment by overcoming some of the most challenging obstacles in current precision medicine.
Accelerating Cancer Control Efforts Through Implementation Science
Research discoveries are the driving force behind every clinical intervention that improves survival and quality of life and every new policy designed to advance public health. However, there is a gap between what we know can improve public health and what gets implemented in everyday life and in clinical practice. This gap creates a substantial impediment to public health. According to NCI, implementation science is a field of research that utilizes scientific approaches to find the best ways to integrate proven, effective interventions into routine health care and public health settings to bridge the gap between evidence and practice. Implementation science is fundamental to cancer control, which is defined as a collective approach aimed at reducing cancer risk, incidence, morbidity, and mortality, and improving quality of life.
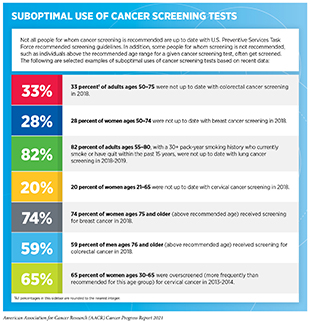
As discussed earlier in the report, there is substantial evidence that HPV vaccination among adolescents can prevent HPV-related cancers and that cancer screening among average-risk individuals can reduce mortality from the screened cancer. However, in the United States, the current uptake of HPV vaccination as well as cancer screening tests is suboptimal among the eligible populations (see sidebar on Suboptimal Use of Cancer Screening Tests). Similarly, there are limitations in the uptake of genetic testing of tumors in clinical practice, even though this is an area with proven health benefits for cancer patients (642)Roberts MC, Kennedy AE, Chambers DA, Khoury MJ. The current state of implementation science in genomic medicine: Opportunities for improvement. Genet Med. 2017;19(8).. Closing the gap between our current knowledge of cancer etiology, prevention, diagnosis, treatment, and survivorship and what is provided as standard care will require significant advances in implementation science. Beyond the United States, implementation science will be key to efforts aimed at reducing the global burden of cancer because it can accelerate cancer control, especially in low- and medium-income countries (LMICs) which are disproportionately affected by cancer.
Researchers are currently assessing implementation of numerous cancer interventions across the continuum of clinical care. Areas of high priority include local adaptation of broad evidence-based interventions, long-term sustenance of effective interventions, advancement of cancer health equity, and policy implementation, as well as de-implementation of harmful or suboptimal practices (e.g., cancer screening among those who may not benefit from screening, such as individuals above the recommended screening age) (643)Oh A, Vinson CA, Chambers DA. Future directions for implementation science at the National Cancer Institute: Implementation Science Centers in Cancer Control. Transl Behav Med. 2021;11(2).. Ongoing investigations in implementation research, many of which are funded by NCI, focus on diverse topics that include increasing cancer screening in underserved communities, enacting tobacco control policies, and improving care of cancer survivors, among others (644)Neta G, Clyne M, Chambers DA. Dissemination and implementation research at the national cancer institute: A review of funded studies (2006-2019) and opportunities to advance the field. Cancer Epidemiol Biomarkers Prev. 2021;30(2).. The overarching goal of these projects is to develop interventions that improve cancer outcomes in both clinical and community settings. To maximize the impact of implementation research on public health, NCI is pursuing a multipronged approach that includes developing methodologies and measures to advance implementation science, increasing access to tools and resources for implementation research (e.g., establishing implementation science laboratories in both health and community settings), disseminating knowledge and data on evidence-based cancer control interventions to all stakeholders, and establishing training programs such as Mentored Training in Dissemination and Implementation in Cancer.
Implementation science was recognized as one of the scientific priority areas by the Cancer Moonshot Blue Ribbon Panel. In September 2019, NCI launched the Implementation Science Centers in Cancer Control initiative to advance this priority. The Blue Ribbon Panel had identified several areas of focused support including: accelerating the delivery of colorectal cancer screening, follow-up, and referrals to care in regions of the United States where screening rates are below national standards; enhancing the delivery of tobacco cessation treatments; and developing approaches to identify and care for individuals with inherited cancer syndromes (645)NIH, National Cancer Institute. Implementation science – opportunities in cancer research. [cited 2021 Aug 9]..
The field of implementation science is ripe with opportunities and holds immense potential for reducing the burden of cancer for all Americans (646)Mitchell SA, Chambers DA. Leveraging implementation science to improve cancer care delivery and patient outcomes. In: Journal of Oncology Practice. Vol 13. ; 2017.. It is hoped that, through coordinated efforts across national, regional, local, and community partners, implementation science can translate knowledge generated from research discoveries into clinical practice and transform the delivery of evidence-based care across the cancer control continuum.